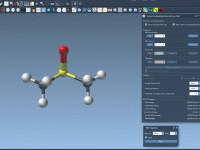
Understanding Property Visibility in SAMSON’s Property Model Attributes
Effortless Control with the ‘visible’ Attribute in SAMSON
Understanding and Using Camera Attributes in Molecular Modeling
Create Molecular Animations Easily with SAMSON’s Animator
For molecular modelers, presenting their work effectively can be challenging. Whether you are summarizing research findings, explaining molecular mechanisms, or preparing visuals for tutorials or publications, animations and presentations can dramatically enhance your storytelling. However, creating professional molecular animations may…
Exploring Topology Flexibility with the Interactive Modeling Universal Force Field
Simplify Molecular Trajectory Exploration with the Play Reverse Path Animation
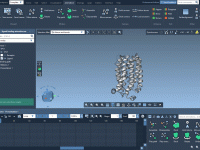
Step-by-step optimization of molecular transition paths with P-NEB.
For molecular modelers, visualizing and optimizing transition paths between molecular conformations can often feel like a tedious puzzle. Paths between states are critical to understanding reaction mechanisms or binding/unbinding processes, especially for systems like protein-ligand complexes. However, crafting and perfecting…
Simplifying Molecular Animations: The Flash Effect Explored
For molecular modelers and researchers, animating molecular structures can provide powerful insights, aid in presentations, or simply enhance visual communication. However, ensuring precise control over what appears and disappears during molecular animations isn’t always straightforward. This is where the Flash…