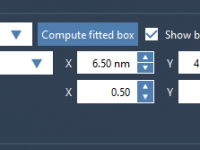
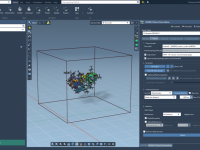
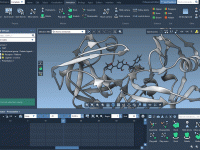
Quickly Identify Side Chains by Name, Charge, Size and More
If you’re modeling biomolecular systems, navigating large molecules to find specific side chains with certain characteristics can be time-consuming. Whether you’re optimizing a binding site, running electrostatics calculations, or preparing a molecule for coarse-grained simulations, filtering side chains by their…
Selecting Atoms Like a Pro in SAMSON: An Introduction to NSL
Stop Searching for Files: Streamline GROMACS Input Selection with Auto-Fill
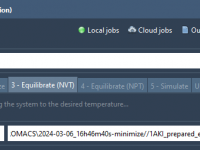
Anyone working with molecular simulations knows the frustration of constantly navigating folders to locate the correct input files—especially when transitioning between simulation steps like energy minimization, NVT equilibration, and NPT equilibration. If you’re using the GROMACS Wizard in SAMSON, there’s…
Effortlessly Identify Hydrogen Bond Donors and Acceptors in Your Molecular Models
When modeling molecular systems, identifying hydrogen bond donors and acceptors quickly and accurately is often crucial. Whether you’re preparing structures for docking, analyzing interactions in a protein-ligand complex, or filtering atoms for visualization, being able to specify which atoms participate…
Quickly Find Specific Chains Using NSL in SAMSON
Avoiding Periodic Pitfalls: Setting Up the Box for COM Pulling in Molecular Simulations
Filtering Aromatic Atoms, the Smart Way
Working with aromatic systems is an essential part of structure-based molecular modeling, from visualizing planar ring systems to performing conformational analysis on drug-like molecules. But selecting these atoms precisely can be time-consuming—especially when your molecules are large or complex. This…