Avoiding Common Mistakes with Bond Detection in Molecular Simulations
Quickly Selecting Visible or Hidden Nodes in SAMSON
Filtering Molecular Structures in SAMSON with NSL: A Simple Yet Powerful Tool
Trying Molecular Modeling for the First Time? These Built-in Tutorials Might Help
Creating Your Own Visual Presets in SAMSON: A Molecular Modeling Shortcut Worth Knowing

If you often find yourself applying the same sequence of visualizations when working on molecular systems—setting colors, choosing representations, focusing on ligands—you are not alone. Consistency in visualization is key for insights, collaboration, and publication. But endlessly repeating the same…
Making Molecular Projects Self-Contained with Embedded Files in SAMSON
Keeping Your Molecular Projects Portable with Embedded Files in SAMSON
A Practical Guide to WHAM-Based PMF Analysis in GROMACS Wizard
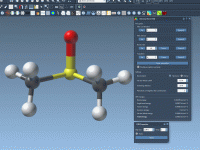
Breaking and Making Bonds while Simulating Molecules with IM-UFF
In traditional molecular modeling workflows, editing structures often means stopping the simulation, changing the topology manually, and restarting the process. This interrupts modeling intuition and introduces friction, especially during rapid prototyping or educational demonstrations. Wouldn’t it be more natural to…