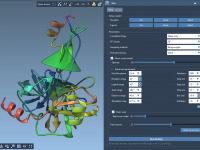
Two Ways to Add Atoms in SAMSON (and When to Use Each)
Controlling the Pace of Molecular Animations with Pause Frames
Struggling With GROMACS Index Groups? Here’s a Simple Way to Manage Them Visually
Creating and managing index groups in GROMACS can be tedious, especially when working with large biomolecular systems. Whether you’re setting up pulling groups, defining restraints, or preparing simulations, it often requires command-line scripts or careful hand-editing of selections. Fortunately, there’s…
Choosing What to Import After Your NVT Equilibration
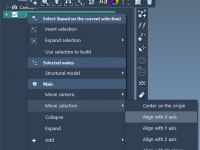
A Quick Way to Align Molecular Structures in SAMSON
Easily Select Residues Based on Charge and Polarity in SAMSON
Working with complex biomolecular structures often requires isolating specific types of residues, such as those that are charged or hydrophobic. This task frequently arises in molecular modeling workflows—whether you’re designing mutations, analyzing ligand binding, or preparing systems for simulations. In…