Still Editing .mdp Files Manually? Try This Instead.
For many molecular modelers using GROMACS, editing .mdp (molecular dynamics parameter) files manually has long been the norm. It’s effective but error-prone, especially when experimenting with different simulation setups like energy minimization, equilibration (NVT, NPT), or production runs. Copy-pasting parameters,…
Get Better NMR Structures Faster: How the FIRE Minimizer Helps
Elevate Your View: How Pedestal Camera Animations Improve Molecular Presentations
Make Atoms Fade Away: A Visual Technique for Clearer Molecular Presentations
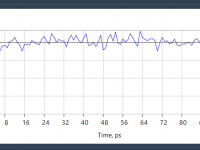
Avoid Wasting Simulation Time: Handling Temperature Stabilization During NVT Equilibration
Which Molecular File Formats Can SAMSON Read and Write?
Need To Share Your Molecular Animation? Here’s How to Export It as a Movie in SAMSON
How to Instantly Clean Up Your Molecular Visuals Using Visual Presets in SAMSON
Creating clear and informative molecular visualizations can quickly become time-consuming—especially when dealing with complex systems like protein-ligand complexes, membrane assemblies, or large macromolecular structures. If you’ve ever found yourself spending more time setting up your visual representations than analyzing your…
Progressively Hiding Atoms in Molecular Animations
When you’re creating molecular animations, there are moments when simply making an atom vanish instantly feels abrupt or visually jarring. Whether you’re teaching, presenting, or designing a publication-ready visual, gradual transitions can significantly improve clarity and storytelling. One frequently encountered…