Reducing Presentation Confusion with the Shown Animation in SAMSON
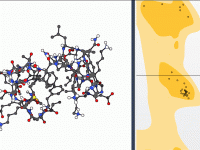
Building Lipid Layers Around Proteins – A Practical Guide for Molecular Modelers
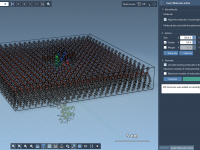
Setting up lipid membranes around proteins is a recurring task in molecular modeling and simulations, particularly for those working on membrane proteins or studying membrane-protein interactions. However, assembling lipid layers manually can be tedious and time-consuming. Fortunately, SAMSON’s Molecular Box…
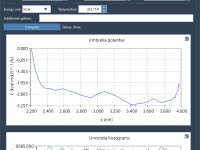
When Reaction Coordinates Aren’t Enough: Visualizing PMF Coverage in GROMACS Wizard
Geometry Optimization Without the Wait: When to Choose FIRE Over Steepest Descent
Quickly Find What’s Visible in Your Molecular Scene with NSL
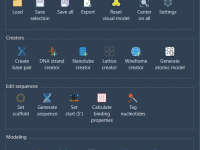
In large-scale molecular simulations, visual clutter is a common issue. Whether you’re analyzing protein-ligand interactions or building complex assemblies, identifying visible elements in your scene is crucial. Fortunately, SAMSON’s Node Specification Language (NSL) includes presentation attributes that make it much…
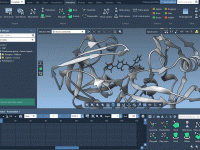
A Clearer Look at Molecules: Visual Models in SAMSON
From Design to Simulation: How to Export DNA Nanostructures with Adenita
Syncing Trajectories with Ease: Using the Play Path Animation in SAMSON
One of the recurring pains in molecular modeling is the clear and synchronized visualization of structural transitions—whether you’re browsing a molecular dynamics trajectory, comparing different conformers, or showing simulation results in a presentation. When you need to animate changes seamlessly…