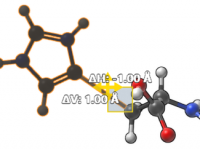
Following Molecules in Motion: How to Make the Camera Track Atoms Automatically in SAMSON
Bringing Molecular Models to Life with Camera Animations in SAMSON
Creating compelling molecular animations is often a challenge for researchers who want to clearly communicate dynamic structural changes, interactions, or spatial relationships between molecules. One of the most common frustrations when preparing presentations, videos, or educational materials is how to…
Creating Attention-Grabbing Molecular Visuals With the Pulse Animation
Sharing Interactive Python Workflows in Molecular Models
Making Elements Appear Naturally in Molecular Presentations
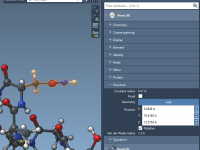
Filtering Molecular Structures in SAMSON: A Closer Look at the Document View and Node Specification Language
When working with complex molecular models, identifying and isolating specific structural elements—like side chains, ligands, or individual residues—can be daunting. This challenge becomes even more critical when models get large, and selections based on visual inspection simply aren’t feasible. SAMSON…