Transforming SMILES Strings into 3D Structures with Ease
For molecular modelers, a common task is transforming SMILES (Simplified Molecular Input Line Entry System) strings into 3D molecular structures for simulations and analysis. However, doing this manually or using cumbersome workflows can become a time-intensive challenge, especially for large-scale…
Integrating Useful Apps into Your Molecular Modeling Workflow
Streamlining Molecular Modeling with File Attributes in SAMSON
Molecular modelers often deal with complex data structures, requiring precise organization and customization to efficiently manage their projects. Whether it’s targeting specific molecular files or managing node-specific details, SAMSON provides powerful tools to simplify these tasks, including its file attributes.…
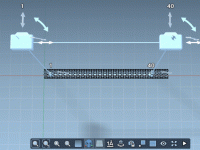
Effortlessly Adjust Horizontal Camera Movements with the Truck Camera Animation in SAMSON
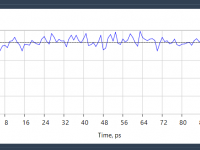
Streamlining Molecular Dynamics: NVT Equilibration Made Simple
Creating Your Custom Importers in SAMSON: A Guide for Molecular Modelers
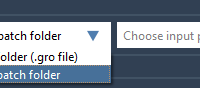
Simplify Your Molecular Simulations with Batch Preparation in SAMSON’s GROMACS Wizard
Choosing the Right Unit Cell for Periodic Boundary Conditions in GROMACS.
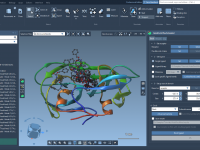
Streamline Molecular System Visualization with SAMSON’s Visual Presets
For molecular modelers, one of the most time-consuming aspects of their work is achieving a clear, visually informative representation of complex molecular systems. Manually configuring visual representations and applying color schemes to large protein-ligand structures can be tedious and error-prone.…