Author: OneAngstrom
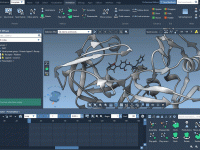
Streamline Molecular Presentations with Pulse Animation in SAMSON
Mastering Segment Attributes: Simplifying Molecular Modeling
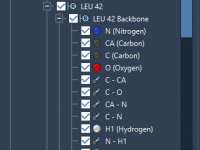
Molecular modelers often face the challenge of efficiently analyzing and managing complex molecular structures. Attributes such as visibility, composition, and charge can provide valuable insights, but working with them can become cumbersome. This is where understanding segment attributes in SAMSON’s…
Understanding Render Preset Attributes in SAMSON
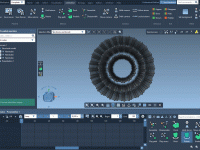
Effortless Simulation in Molecular Design: Adding Keyframe Animations in SAMSON.
Understanding Note Attributes in SAMSON’s NSL
Simplifying Molecular Rotations Using Geometry-Based Animations
How Shared Documents Streamline Collaboration in Molecular Modeling.
Mastering Molecular Organization in SAMSON: An Overview of the Document View
Understanding Segment Attributes in Molecular Modeling
If you’re involved in molecular modeling, you understand the challenge of working with large-scale molecular structures and navigating their complexity. SAMSON’s Node Specification Language (NSL) provides powerful tools to solve these challenges, allowing precise interaction with structural elements like segments.…