Streamline Your PMF Analysis with GROMACS Wizard’s WHAM Tool
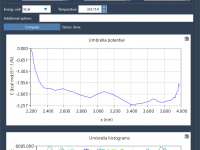
For molecular modelers engaged in understanding reaction-coordinate dynamics, generating accurate Potential of Mean Force (PMF) profiles is a critical yet often challenging part of umbrella sampling workflows. Fortunately, SAMSON’s GROMACS Wizard provides a structured and user-friendly approach for PMF analysis…