Mastering the Dolly Camera Animation in Molecular Modeling
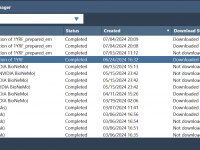
Simplify Cloud-Based Molecular Simulations with SAMSON’s Job Manager
Mastering the ‘Disappear’ Animation for Molecular Models: A Step-by-Step Guide
One common challenge for molecular modelers is finding ways to effectively communicate complex molecular data in presentations and animations. Especially when working with intricate structures, such as molecular assemblies or visualizing dynamic processes, the ability to gradually highlight relevant features…
Creating a Seamless ‘Disassemble’ Effect with SAMSON Animator
Streamline Molecular Modeling with Dock Animation in SAMSON
Streamlining Molecular Interactions with Dock Animation in SAMSON
Decoding Protein Structures with AlphaFold-2 on SAMSON
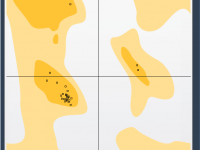
Master Protein Conformations with the Interactive Ramachandran Plot.
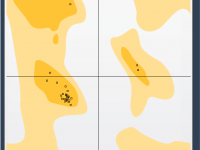
Validating and refining protein structures can be a challenging task for molecular modelers. Modeling accuracy directly impacts the success of simulations, predictions, and experimental designs. One particularly useful tool to address these challenges is the Interactive Ramachandran Plot provided by…