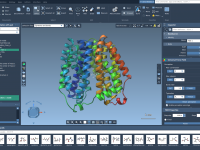
A Practical Way to Explore Molecular Substitutions with SMARTS Patterns
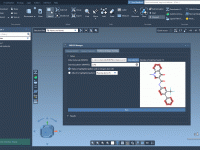
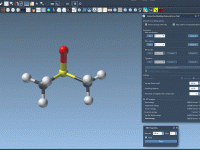
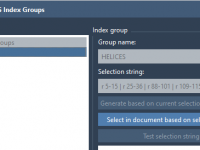
Exploring molecular analogs is a common challenge in molecular modeling. Whether you’re designing compounds for drug discovery, studying structure-activity relationships, or preparing structures for docking, generating analog series efficiently is essential. Fortunately, the SMILES Manager in SAMSON provides a straightforward…