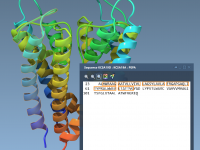
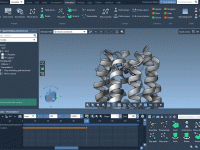
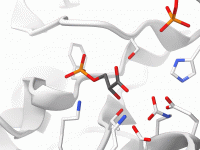
Enhance Molecular Depth Perception with Ambient Occlusion in SAMSON
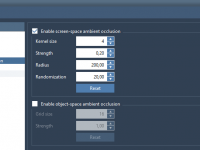
For molecular modelers and designers, depth perception is essential to understanding complex structures. Without proper rendering techniques, 3D models can appear flat, making it difficult to grasp critical spatial relationships within molecular systems. Thankfully, SAMSON offers a powerful tool to…