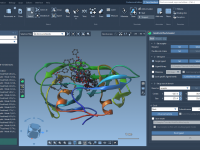
Transform Your Molecular Modeling Workflow with IM-UFF
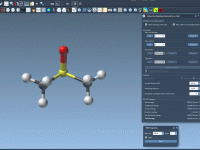
For molecular modelers, one common challenge is efficiently building, editing, and simulating molecular structures that may undergo significant topological changes. SAMSON’s Interactive Modeling Universal Force Field (IM-UFF) offers a solution tailored to this need, seamlessly blending interactive molecular modeling with…
Redefining Molecular Modeling Efficiency with SAMSON’s Interaction Designer
Getting Familiar with the SAMSON Interface: A Guide for Molecular Modelers
Mastering Pause Animations in Your Molecular Presentations
Mastering Sequence Views to Simplify Molecular Modeling
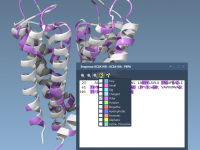
For molecular modelers, managing complex molecular structures with multiple residues and chains can often feel overwhelming. Keeping track of sequences and their corresponding properties across different views commonly leads to frustration when trying to align selections or visualize modifications. This…
Fine-Tuning Molecular Structures: Minimizing Specific Components in SAMSON
For molecular modelers, minimizing structures to achieve optimized geometries is a routine task. However, selectively minimizing only specific parts of a molecular system can sometimes be challenging, especially when working with large or complex structures. Thankfully, SAMSON provides features tailored…
Simplify Molecular Visualizations with Visual Presets in SAMSON
One of the frequent challenges for molecular modelers is quickly and effectively visualizing complex molecular systems. These systems often involve various components like ligands, receptors, and structural elements, each requiring distinct visual representations and color schemes for clear understanding. Instead…