Controlling Protein Motion: A Practical Guide to the Sampling Box in SAMSON’s Protein Path Finder
Choosing the right discrete color palette in SAMSON
How to Set Up Energy Evaluation Models for Ligand Pathway Analysis in SAMSON
Interpolating Conformational Changes: How to Visualize Macromolecular Transitions in SAMSON
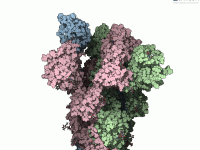
In molecular modeling, creating smooth and accurate visualizations of molecular transitions is often a challenge. Whether you’re investigating protein-ligand interactions or large conformational changes in biomolecular complexes, understanding the path a molecule takes between two states can be essential for…
Making Atoms Dance: A Simple Way to Create Molecular Motion
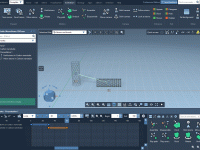
Creating animations of molecular structures is not just about aesthetics—it’s a powerful way to communicate behavior, mechanisms, and design insights. But for many molecular modelers, translating molecular motions into effective animations can be time-consuming and unintuitive. One frequent challenge: how…
Switching Selections in Seconds: A Look at Quick Groups in SAMSON
Translating Molecules with AI-Generated Python in SAMSON
Pausing Molecular Presentations Effectively in SAMSON
Creating molecular animations can be a powerful way to communicate complex structural transitions, conformational changes, or step-by-step molecular mechanisms. However, when presenting these animations, it’s often necessary to slow down or emphasize a particular moment in the animation. For instance,…