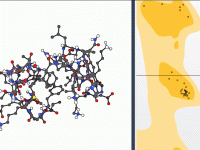
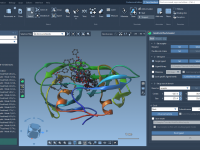
Creating and Breaking Bonds in Real Time with IM-UFF
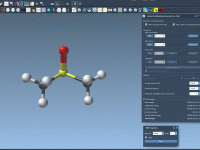
One common challenge for molecular modelers is editing molecular structures while maintaining physical realism. Whether you’re designing molecules, studying reaction pathways, or exploring hypothetical structures, you often need to dynamically add or break bonds. Doing this manually can be time-consuming…