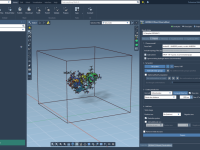
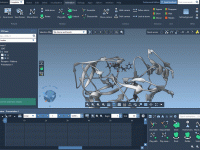
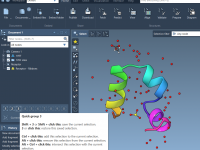
Quickly Show or Hide Labels in SAMSON Using NSL
Quickly Find the Files You Need in Complex Molecular Projects
How to Add and Delete Atoms During a UFF Simulation in SAMSON
When Whole-Protein Alignment Doesn’t Cut It: Superimposing Specific Regions in SAMSON
Avoiding Water Clashes in Coarse-Grained MD
Installing Molecular Modeling Software Without Admin Rights: Here’s How
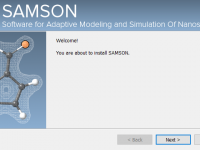
Many molecular modelers—especially students and researchers in institutional environments—face a recurring and frustrating challenge: installing advanced software tools on machines where they don’t have administrative privileges. If you’ve ever been excited to try a new modeling platform, only to be…