A Clearer View of Protein-Ligand Interactions: 2D Meets 3D
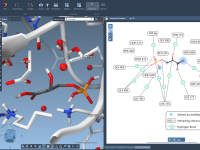
When analyzing protein-ligand complexes, understanding non-covalent interactions—like hydrogen bonds, van der Waals contacts, and pi-stacking—can feel overwhelming in 3D alone. That’s where 2D interaction diagrams make a difference: they simplify complexity and help researchers interpret molecular details more clearly. Interaction…