Making Molecules Appear at Just the Right Moment
Visual Transitions in Molecular Presentations: Adding Background Interpolations
Tired of Manual Plugins? Here’s How SAMSON Makes Extension Management Easier
For molecular modelers, customizing software to suit specific workflows often means managing a jungle of plugins and add-ons. Whether it’s downloading, installing, updating, or simply keeping track of versions—plugin management can be time-consuming and error-prone. SAMSON provides a different solution…
Quickly Generate Carbon Nanotube Models with Precision (No Scripting Needed)
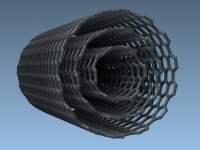

For many molecular designers and materials scientists, building accurate models of carbon nanotubes (CNTs) often represents a bottleneck in their workflow. Whether you’re simulating transport phenomena, designing sensors, or exploring nanostructured materials, it’s crucial to start with well-defined geometries. But…
Interpolating Molecular Motions: How to Model Spike Transitions Without Guesswork
When you are studying large macromolecular systems, one notoriously difficult task is to simulate plausible transitions between two known conformations. Whether you’re investigating protein folding, loop dynamics, or viral mechanisms, modeling intermediate structures between an open and closed state can…
A Simple Way to Define Custom Index Groups for GROMACS Simulations
Quickly Isolate Property Models Based on Visibility and Selection Flags
Easily Find and Filter Notes in Complex Molecular Models
When working with detailed molecular models in SAMSON, it’s easy for your workspace to become difficult to navigate. Notes scattered throughout models often contain crucial information—observations, temporary labels, or structural annotations—but they can quickly become visually overwhelming. Thankfully, with the…