Mastering Label Visibility and Customization in SAMSON
Simplifying NPT Equilibration with the GROMACS Wizard
Exploring Camera Node Attributes in SAMSON
How the ‘Stop’ Animation Effect Helps Organize Molecular Models
For molecular modelers looking to create clear and professional presentations of their work, organizing animations into well-defined segments or “slides” can be a challenge. This challenge becomes particularly relevant for those showcasing complex molecular dynamics, reactions, and processes. Fortunately, SAMSON’s…
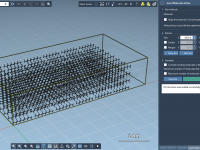
Effortless Molecular Box Generation with SAMSON’s Molecular Box Builder
Step-by-Step Guide to Using the Appear Animation in Molecular Modeling
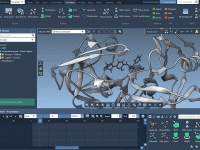
Visualizing molecular structures with precision is key for molecular modelers who strive to communicate ideas effectively, simulate processes, or present their research. One common challenge is progressively making elements in your model visually appear, particularly when dealing with transparency-sensitive structures…
Simplifying Molecular File Imports in SAMSON: A Quick Guide
Easily Reset Attributes to Default in SAMSON’s Inspector
Simplify Molecular Visualizations with the Rotate Animation in SAMSON
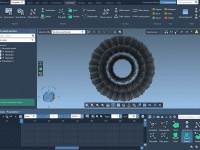
Efficient visualization and understanding of molecular behavior is paramount for molecular modelers. Whether you’re presenting findings or analyzing molecular dynamics, clear and controlled animations can make a significant difference. One powerful yet often underutilized feature in SAMSON is the Rotate…