Streamline Molecular Loading in SAMSON
A Stress-Free Guide to Installing and Starting SAMSON
Freezing Atoms for Precise Molecular Minimization in SAMSON.
Understanding Structural Group Attributes in SAMSON’s Node Specification Language.
Mastering the Move Camera Animation in Molecular Presentations
Streamline Protein Sequence Alignment with SAMSON’s Protein Aligner.
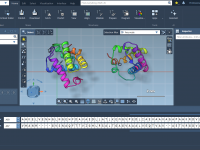
Protein sequence alignment is a critical step for molecular modelers solving a variety of structural and functional questions. However, aligning sequences and comparing structures often involve juggling between different tools and formats, which can be time-consuming and error-prone. SAMSON’s Protein…
Exploring Animation Effects in SAMSON: Bring Molecular Structures to Life
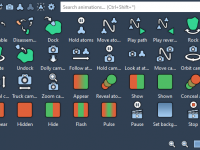
As a molecular modeler, presenting your work effectively is just as important as the technical rigor of simulations. Whether you’re demonstrating molecular docking, pathways, or structural transformations, animation can help you communicate complex information persuasively and clearly. Fortunately, SAMSON –…
Streamline Residue Selection and Visualization with Sequence View
Effortlessly Manage SAMSON Extensions for Your Molecular Modeling Needs.

For molecular modelers who want to customize and expand their software capabilities, managing extensions can sometimes feel overwhelming. SAMSON makes this process significantly easier with its Marketplace and built-in extension management tools. Here’s a step-by-step guide to understand and maximize…