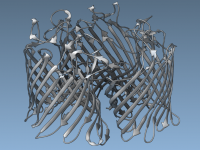
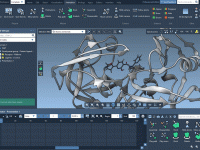
Achieve Stunning Depth Perception with Ambient Occlusion in Molecular Models
Mastering Bond Length Queries in Molecular Modeling
Simplify Your Molecular Simulations with the ‘Hidden’ Animation
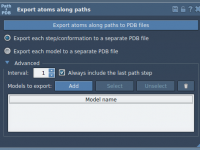
Export Specific Atom Trajectories With SAMSON for Advanced Molecular Modeling.
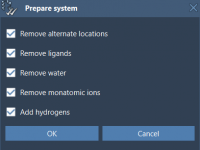
Why Validating Protein Structures Before Simulations Saves Time and Effort
Mastering Constrained Molecule Simulations with SAMSON’s Simulate Animation
Simulating molecular systems plays a pivotal role in understanding and designing nanoscale systems. However, molecular modelers often encounter challenges when trying to integrate constraints, such as fixed atomic positions or specific configurations, into their simulations. Luckily, SAMSON’s Simulate animation offers…
Optimizing Protein Simulations with Sampling Boxes in Protein Path Finder.
Molecular modelers seeking highly efficient ways to simulate protein motions often encounter a common challenge: constraining the conformational exploration within biologically relevant regions. Overlooking such constraints can lead to time-consuming computations and irrelevant conformations. A practical solution to this problem…
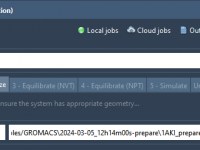
Streamlining Energy Minimization: How to Select Your Input Structure in GROMACS Wizard
Simplifying Initial Conformations for Umbrella Sampling in GROMACS Wizard
Molecular modelers often face challenges when setting up Umbrella Sampling simulations, particularly in obtaining appropriate initial conformations. If improperly prepared, this crucial step can complicate the entire workflow and lead to inefficiencies in analyzing the Potential of Mean Force (PMF).…