Author: OneAngstrom
Effortless Residue Selection and Coloring with Sequence Views in SAMSON
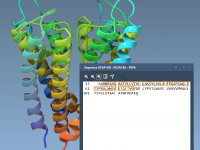
For molecular modelers, analyzing and manipulating biomolecular structures across 1D, 2D, and 3D representations is often a daunting task. Switching between these views while maintaining coherence can feel cumbersome. Fortunately, SAMSON simplifies this process through its interactive Sequence View feature,…
Mastering Folder Attributes in SAMSON’s Node Specification Language
Molecular modeling often requires working with large datasets and intricate structures nested within folder-like organizational frameworks. Navigating and querying such data precisely can be overwhelming without the right tools. SAMSON’s Node Specification Language (NSL) offers a well-designed system of folder…