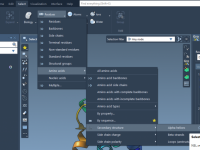
Selecting Visual Presets in SAMSON: A Quick Guide with the NSL
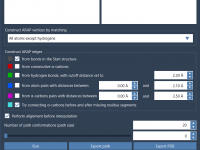
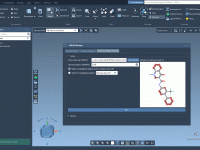
When working on molecular models in SAMSON, visualization is often more than just aesthetics—it’s essential. Different render presets can highlight atomic structures, interaction networks, or molecular surfaces, which all help communicate scientific insight effectively. But when projects become complex, selecting…