Mastering Interactive Simulations in SAMSON: A Step-by-Step Guide.
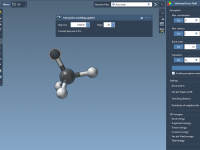
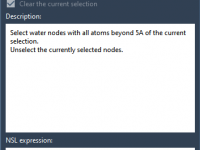
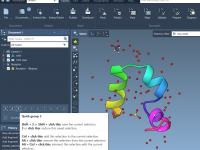
For molecular modelers, achieving accurate atomic behaviors and understanding molecular dynamics can be a challenge. Interactive simulations provide a way to experiment and control simulations while maintaining flexibility, and SAMSON offers powerful tools to implement this effectively. This blog post…
Mastering Custom Color Palettes in SAMSON
Exploring Animation Effects in Molecular Modeling
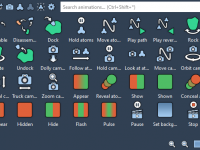
Creating animations can greatly enhance molecular modeling workflows, making complex presentations and analyses more effective. Animations allow modelers to visualize molecular processes dynamically, communicate results more vividly, and even generate instructional materials for teaching and knowledge sharing. SAMSON offers an…