Unveiling Biomolecular Dynamics with SAMSON’s Normal Modes Advanced Module
Simplify Molecular Modeling with the Pulse Animation
Simplify Molecular Visualization with Color Palette Reversal
Simplifying Protein Preparation in SAMSON for Transition Pathway Analysis
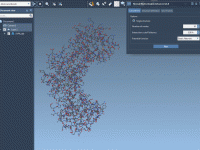
When modeling protein conformational transitions, setting up your system correctly is crucial to ensure accurate and physically meaningful results. However, for many molecular modelers, the preparation of protein systems—removing unwanted elements, fixing missing residues, and ensuring hydrogen atoms are correctly…
Leveraging Folder Attributes for Molecular Modeling Efficiency.
As a molecular modeler, organizing and manipulating complex molecular systems efficiently can be challenging. Key to addressing this challenge is understanding and leveraging folder attributes in SAMSON’s Node Specification Language (NSL). Folder attributes offer a streamlined way to access, filter,…
Simplifying Molecular Visualization with Visual Presets in SAMSON
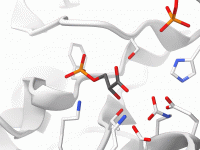
For many molecular modelers, one of the recurring challenges is efficiently visualizing complex molecular systems. The amount of time spent meticulously adjusting representations, selecting color schemes, and ensuring clarity can quickly become overwhelming—especially when there are deadlines or presentations just…
Optimizing Molecular Structures in SAMSON: A Guide to Interactive Minimization.
Understanding Backbone Node Attributes in Molecular Modeling
Efficiently Managing Computational Jobs in SAMSON Cloud.
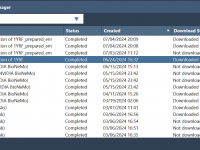
For molecular modelers, working on complex molecular dynamics or protein structure prediction tasks often involves computationally intensive calculations. Managing these tasks effectively can be a challenge, especially when dealing with cloud-based workflows. This is where the Job Manager in SAMSON…