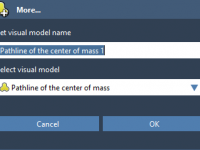
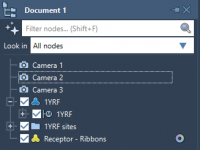
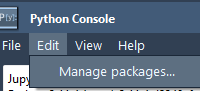
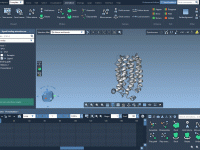
A Simpler Way to Add, Try, or Remove a SAMSON Extension

As molecular modelers, we often need specific tools for our research—whether it’s visualization tweaks, structure preparation helpers, or access to advanced simulation algorithms. But installing and managing plug-ins or modules across platforms can be frustrating and time-consuming, especially when you…