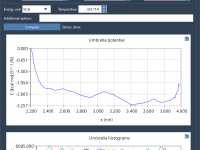
A Practical Guide to WHAM-Based PMF Analysis in GROMACS Wizard
Breaking and Making Bonds while Simulating Molecules with IM-UFF
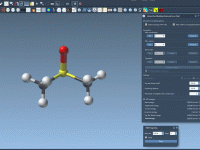
In traditional molecular modeling workflows, editing structures often means stopping the simulation, changing the topology manually, and restarting the process. This interrupts modeling intuition and introduces friction, especially during rapid prototyping or educational demonstrations. Wouldn’t it be more natural to…
Filtering Molecular Bonds by Type in NSL
No Admin Rights? No Problem: Installing SAMSON Seamlessly
Building Lipid Bilayers Around Proteins in SAMSON
Choosing the Right Color Palette in Molecular Modeling: A Guide to Discrete Palettes in SAMSON
Quickly Filter and Control Light Nodes in SAMSON Using the NSL
Swapping Atoms, Saving Time: Quick Analogue Generation in SAMSON
FASTA In, Structure Out: Predicting Proteins with AlphaFold-2 in SAMSON
Predicting protein structures used to be a long, resource-heavy process, often requiring local installations or complex data preparation. Today, especially with the introduction of AlphaFold-2, molecular modelers have access to powerful prediction engines remotely. However, integrating these capabilities into real…