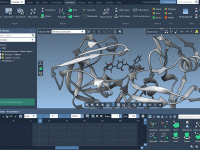
Simplifying Transition Path Optimization with P-NEB in SAMSON
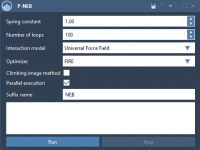
Molecular modeling often requires identifying precise transition paths between states of a system, such as ligand binding and unbinding or conformational shifts in proteins. These paths help researchers understand key mechanisms, but generating accurate paths can be computationally challenging. Enter…
Easily Switch to Your Custom GROMACS Version in SAMSON
Mastering Solvent Options for Coarse-Grained Simulations in GROMACS Wizard
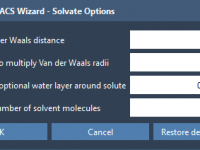
When preparing coarse-grained (CG) molecular systems for simulations, ensuring proper solvation can be one of the trickiest yet most critical steps. Solvent configuration directly impacts the accuracy and stability of molecular dynamics (MD) simulations, especially for coarse-grained models like those…
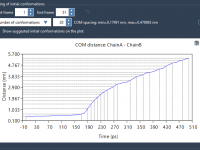
Exporting Atom Trajectories Along Paths: A Step-by-Step Guide
Molecular modelers often face the challenge of exporting atomic trajectories to analyze reaction coordinates, evaluate free energy changes, or visualize molecular pathways. SAMSON’s Export Along Paths extension simplifies this process, enabling users to export atomic coordinates along defined paths. This…
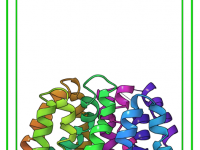
Setting up the Sampling Box in Ligand Pathway Exploration: Why Precision Matters
Effortlessly Manage Molecular Jobs with SAMSON’s Job Manager
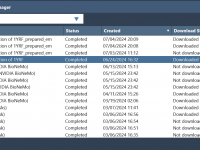
One of the most time-consuming challenges molecular modelers face is managing complex computational workflows. Whether it’s protein structure prediction, molecular dynamics simulations, or advanced tasks like utilizing NVIDIA BioNeMo, staying on top of your calculations can be daunting. SAMSON offers…
Enhancing Molecular Modeling with AlphaFold-2 in SAMSON
Mastering File Formats in SAMSON for Molecular Modeling
Mastering Node Attributes in SAMSON: A Guide for Molecular Modelers
Understanding and effectively utilizing node attributes in SAMSON’s Node Specification Language (NSL) can make a significant difference in how molecular modelers interact with structural data. Whether you’re selecting residues, visualizing specific groups, or focusing on locked nodes, node attributes enable…