Author: OneAngstrom
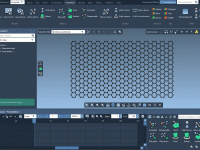
Building Carbon Nanotubes with Intuitive Viewport Interaction in SAMSON
A Quick Dive into Managing Camera Attributes in SAMSON
Practical Guide to Filtering Side Chains in Your Molecular Models
Unveiling Folder Attributes for Molecular Organization in SAMSON
For molecular modelers, efficiently working with complex molecular structures often means managing folders within their design platforms. SAMSON’s Node Specification Language (NSL) offers a powerful set of folder attributes that make molecular organization, querying, and filtering smoother and more precise.…
Streamline Molecular Design with Python in SAMSON
Mastering the Art of ‘Conceal Atoms’ for Molecular Animations
Streamlining Molecular Trajectories: Reverse Path Animation in SAMSON
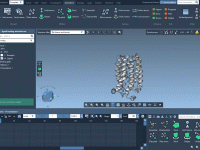
Molecular modeling often involves studying and interpreting dynamic processes, such as conformational changes, binding events, or even complex molecular pathways. One critical challenge many molecular modelers face is efficiently exploring and presenting complex molecular trajectories. The Play reverse path animation…