Author: OneAngstrom
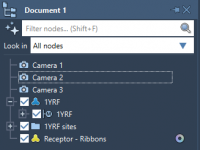
Efficiently Managing Camera Nodes in SAMSON with NSL
Molecular modeling often involves the generation of complex visual representations. Controlling and customizing camera nodes in a molecular modeling environment can significantly enhance these visualizations. In SAMSON, the Node Specification Language (NSL) enables targeted management of camera attributes, streamlining the…
Mastering Camera Handling in SAMSON for Seamless Molecular Modeling.
Understanding Folder Attributes: A Helpful Guide for Molecular Modelers
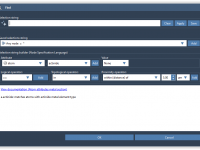
Mastering Molecular Selections with the Find Command in SAMSON
Bring Molecular Models to Life with the Pulse Animation
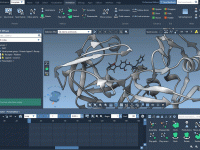
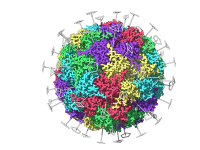
For molecular modelers, effectively visualizing complex structures is essential, especially when presenting to teams or colleagues. A common challenge? Making dense molecular scenes more comprehensible and engaging without overwhelming the audience. Enter SAMSON’s Pulse animation. The Pulse animation is designed…
Effortless Protein Transition Paths with ARAP Interpolation
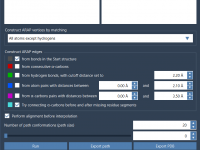
For molecular modelers working with protein dynamics, understanding how a protein transitions between two conformations is often crucial. Yet, generating smooth and biologically meaningful transition paths between these states can be both time-consuming and complex. This is precisely where SAMSON’s…
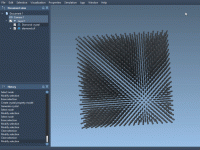
Understanding Defects in Diamond Crystals with SAMSON.
Effortlessly Running Molecular Simulations in the Cloud with GROMACS Wizard.
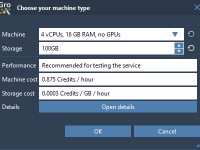
Running molecular dynamics simulations can be resource-intensive, especially when dealing with complex systems or long simulation times. As a molecular modeler, you might have encountered performance bottlenecks or limited hardware on your local machine, hindering experimentation and workflows. Thankfully, the…