Author: OneAngstrom
Master Molecular Modeling with Simple Script Extension in SAMSON
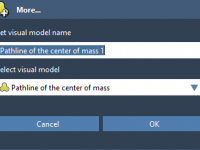
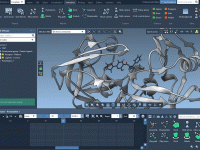
How to Track Molecular Motion with Pathlines in SAMSON
Easily Build a Distinctive Public Profile on SAMSON Connect
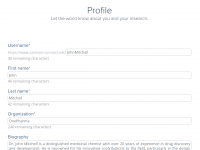
For molecular modelers, establishing a professional presence and communicating effectively with collaborators can be a challenge. The solution? SAMSON Connect’s advanced profile creation tools, which empower users to create detailed, attractive public profiles to share their expertise and build their…
Understanding SAMSON Extension Compatibility: What Every Molecular Modeler Should Know
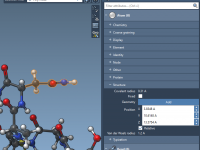
Exploring Camera Attributes for Smarter Molecular Modeling
In molecular modeling and visualization, controlling the perspective and focus of your workspace is essential to gain meaningful insights and clarity. SAMSON’s integrative molecular design platform offers a powerful way to manage perspectives—through camera attributes. If you’ve ever struggled with…