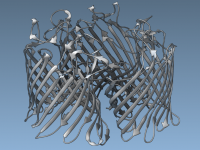
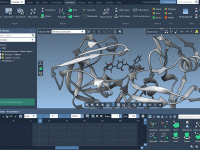
Enhance Molecular Depth with Ambient Occlusion in SAMSON
Master Vertical Movement with the Pedestal Camera in SAMSON
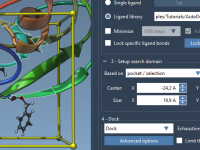
Fine-Tuning Your Docking Search Domain in AutoDock Vina Extended
Simplify Molecular Modeling: Setting Up Your System in SAMSON’s Ligand Path Finder
Simplifying PMF Computation with the GROMACS Wizard
Exploring Symmetry in Protein Design: Generating Replicas Made Simple
Understanding Property Model Attributes in Molecular Design
Making Molecular Visualizations Gradually Come to Life with the Appear Animation
Simplifying Transition Path Optimization with P-NEB in SAMSON
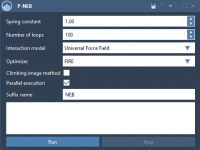
Molecular modeling often requires identifying precise transition paths between states of a system, such as ligand binding and unbinding or conformational shifts in proteins. These paths help researchers understand key mechanisms, but generating accurate paths can be computationally challenging. Enter…