Effortlessly Control Molecular Visibility with the Flash Animation.
How to Add, Try, and Manage SAMSON Extensions Efficiently
Optimizing Ligand Pathways with the Right Sampling Box Setup
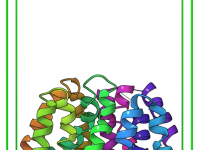
In molecular modeling, identifying ligand unbinding pathways is an essential step in understanding molecular interactions. However, improper configuration of simulation parameters can lead to misaligned results or under-optimized pathways. One highly impactful yet often understated configuration is defining the sampling…
Mastering Molecular Animations: Conceal Atoms Effect Explained
Efficiently Manage Views with Cameras in SAMSON
Enhance Molecular Depth Perception with Ambient Occlusion in SAMSON
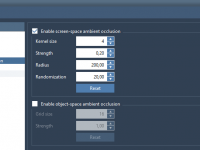
For molecular modelers and designers, depth perception is essential to understanding complex structures. Without proper rendering techniques, 3D models can appear flat, making it difficult to grasp critical spatial relationships within molecular systems. Thankfully, SAMSON offers a powerful tool to…
Synchronizing Sequence and 3D Views in SAMSON: A Practical Guide
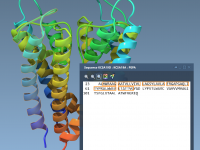
For molecular modelers, managing complex molecular structures across different representations can be a daunting task. A common pain point is ensuring consistent selections between sequence views and 3D visualizations. SAMSON’s Sequence View offers an elegant solution, providing seamless synchronization between…