Mastering Precision: Moving Objects with Snapping in SAMSON
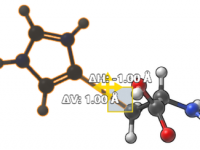
For molecular modelers, positioning structures with accuracy can feel like a delicate balancing act. Whether you’re aligning molecules or arranging complex assemblies, precise movements are crucial to achieving reproducible, high-quality designs. Thanks to SAMSON’s snapping functionality, you can efficiently move…
Mastering Keyframes in Molecular Animations with SAMSON
Enhancing Molecular Visualizations with the Pulse Animation
Creating insightful molecular visualizations often means finding ways to highlight dynamic behaviors or structural emphasis. For molecular modelers, one common challenge is effectively presenting how certain parts of a molecule interact dynamically while managing visual clutter. If you’ve faced this…
Simplify Molecular Animations with Keyframes in SAMSON
Effortlessly Refine Molecular Transition Paths with the P-NEB App
Molecular modelers often face the challenge of identifying accurate transition pathways between states, such as ligand binding or unbinding events. However, ensuring these paths accurately reflect minimum energy trajectories requires sophisticated tools. That’s where SAMSON’s Parallel Nudged Elastic Band (P-NEB)…