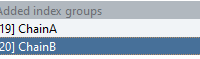
Understanding Node Group Attributes in SAMSON.
Simplifying Molecular Modeling Tasks with SAMSON's Built-in Interactive Tutorials
Mastering the Rock Animation in Molecular Modeling
Enhance Your Molecular Visualizations with Default Color Palettes in SAMSON
Streamline Molecular Interactions with SAMSON’s Editors

As a molecular modeler, you often face the challenge of efficiently manipulating and editing molecular structures—whether you’re generating new models, tweaking geometries, or applying specific transformations. This can be a labor-intensive process, requiring precision and the right tools. But what…
Choosing Unit Cell Shapes in Molecular Simulations: Tips for Efficiency
Step-by-Step Guide to Setting Up a COM Pulling Simulation in GROMACS
Master Horizontal Camera Movements with the Truck Camera Animation
Smoothly Animate Paths in Reverse with SAMSON
When working on molecular modeling, the ability to visualize transitions, trajectories, or conformations between molecular structures is critical. One of the most common challenges is efficiently visualizing reverse dynamics, whether it’s playing a trajectory backward or cycling through conformations. With…