Optimizing Simulation Workflows Using the Simulate Animation in SAMSON
Modern molecular design often involves simulating nanosystems to study their behavior and optimize their structure. Whether you are modeling protein interactions, designing nanomachines, or exploring dynamic molecular systems, running simulations efficiently is crucial to achieving accurate and actionable results. SAMSON’s…
Understanding Node Group Attributes in SAMSON.
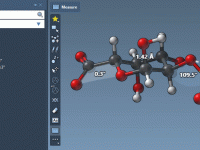
Mastering Measurement Labels in Molecular Modeling with SAMSON
Simplifying Molecular Visualization with the Flash Animation in SAMSON.
Mastering Label Visibility in Molecular Models.

When designing molecular models, managing label visibility efficiently can significantly impact the clarity and presentation of your work. Labels help depict node names, group descriptions, or measurements in the Viewport. However, as your model complexity increases, maintaining label readability becomes…
Simplify Molecular Presentations with the Pause Animation Effect
Simplifying Ligand Setup for Docking with AutoDock Vina Extended
Focused Minimization: How to Minimize a Single Molecule in SAMSON
Streamlining Molecular Modeling with Backbone Attributes in SAMSON
One of the recurring challenges molecular modelers face is effectively managing and analyzing complex molecular backbones in their systems. Whether it’s adjusting visibility for a cleaner workspace, identifying specific attributes, or filtering based on predefined criteria, working efficiently with large…